from pylgs.all import *Polychromatic LGS
A multi-level sodium system excited by 330 nm light
Define a LGS system pumped by 330 nm light:
lgs = LGSSystem(
'Na330',
{
'IntensitySI1': 5000.,
'BFieldG': 0.5,
'EllipticityDegrees1': 45.,
'PolarizationAngleDegrees1': 0,
'DetuningHz1': 1.0832e9,
'LaserWidthHz1': 10.0e6,
'MagneticZenithDegrees': 45.,
'MagneticAzimuthDegrees': 45.,
'SDampingCollisionRatePerS': 4081.63,
'BeamTransitRatePerS': 131.944,
'VccRatePerS': 28571.,
'TemperatureK': 185.,
'RecoilParameter': 1.
}
)The system is made up of 774 density-matrix elements describing 7 atomic levels:
lgs.level_population.rangeLevel(7)
<xarray.DataArray 'XarrayVectorSpace' (Level: 7)> Size: 56B array([0., 0., 0., 0., 0., 0., 0.]) Coordinates: * Level (Level) <U16 448B '3D<sub>3/2</sub>' ... '3P<sub>1/2</sub>'
HTML(', '.join(lgs.level_population.range.coords['Level'].data))
3D3/2, 4P3/2, 3D5/2, 3P3/2, 4S1/2, 3S1/2, 3P1/2
Steady-state model
Adaptively refine velocity groups
Build a steady-state model with adaptively refined velocity groups:
model = lgs.adaptive_stationary_model({}, max_weight=0.02)Solve for the density matrix. There are 77 velocity groups in the model:
sol = model.solve()
sol∅ ⛒ Atomic velocity(77) ⛒ Density matrix(774)
<xarray.DataArray (Atomic velocity: 77, Density matrix: 774)> Size: 477kB
array([[ 4.81176178e-04, 5.34383694e-05, -5.34345398e-05, ...,
-7.62458011e-08, -1.59213581e-10, 4.80993663e-10],
[ 1.57798955e-02, 1.75251498e-03, -1.75236074e-03, ...,
-4.16181537e-06, -1.44907597e-08, 3.47742328e-08],
[ 1.36170875e-02, 1.51285607e-03, -1.51264792e-03, ...,
-6.12657935e-06, -3.65981137e-08, 7.93941184e-08],
...,
[ 3.46659562e-04, 4.29832767e-05, -4.25355464e-05, ...,
1.13082229e-07, -3.42890749e-10, 8.25194594e-10],
[ 8.73280729e-04, 1.00106012e-04, -9.98470213e-05, ...,
2.02082796e-07, -5.61752597e-10, 1.43885993e-09],
[ 4.79681482e-04, 5.35790217e-05, -5.35574066e-05, ...,
7.67512540e-08, -1.60052771e-10, 4.82713824e-10]],
shape=(77, 774))
Coordinates:
* Atomic velocity (Atomic velocity) float64 616B -2.5 -1.5 ... 1.875 2.5
* Density matrix (Density matrix) <U68 211kB 'ρ<sub>Re, (3S<sub>1/2</sub>...Plot the total return flux for each transition as a bar chart:
model.total_flux(sol).visualize()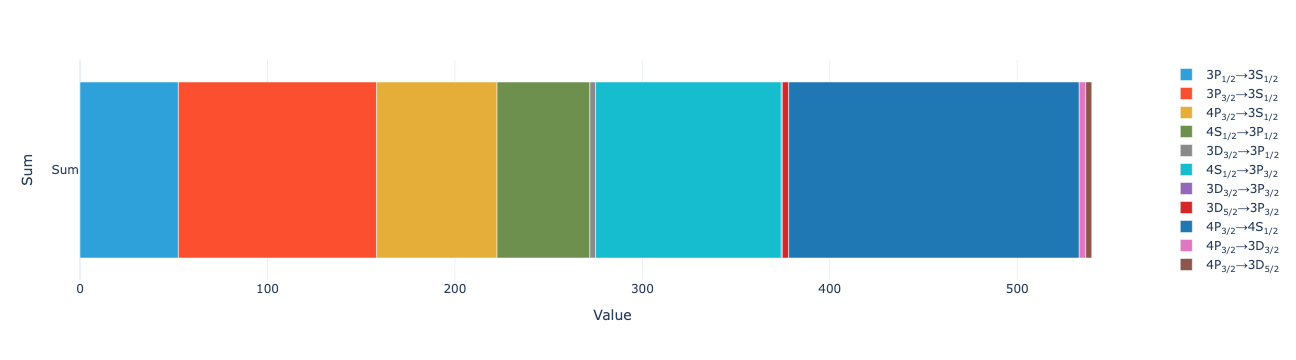
Plot the flux as a function of velocity:
model.flux_distribution(sol).visualize(xaxis_range=(-1, 2))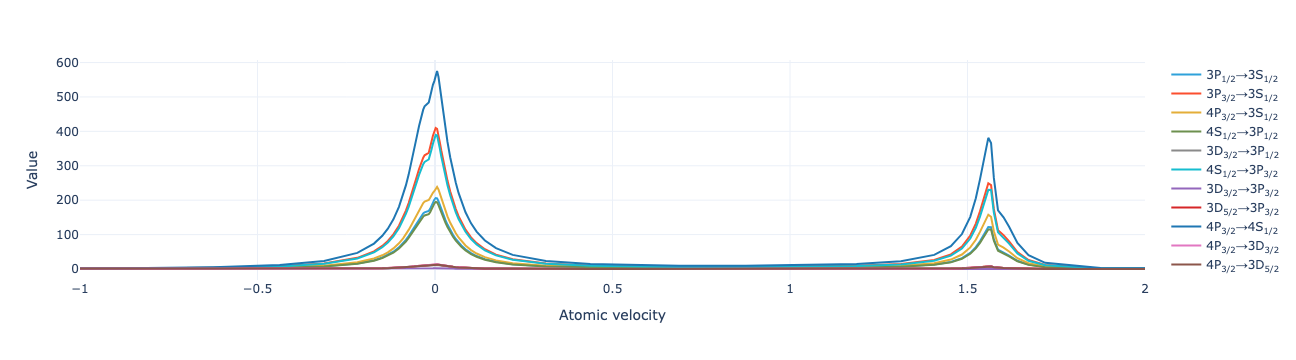
model.flux_distribution(sol).visualize(xaxis_range=(-1, 2))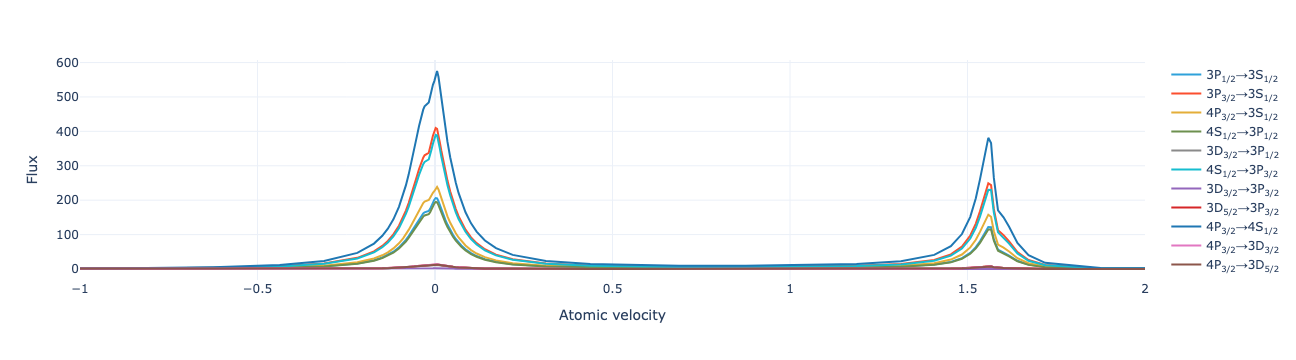
Transient dynamics model
Build a model for the transient dynamics:
model = lgs.instationary_model(
vg=model.data['velocity_groups'],
T=1e-6,
num_values=101
)Set tolerances for the BDF solver:
pymor.basic.set_defaults({
'pylgs.pymor.timestepping.cvode_solver_options.cvode_bdf_atol': 1e-6,
'pylgs.pymor.timestepping.cvode_solver_options.cvode_bdf_rtol': 1e-5,
})Solve for the density matrix as a function of time:
sol = model.solve()solTime(48) ⛒ Atomic velocity(77) ⛒ Density matrix(774)
<xarray.DataArray (Time: 48, Atomic velocity: 77, Density matrix: 774)> Size: 23MB
array([[[ 2.90977780e-04, 0.00000000e+00, 0.00000000e+00, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00],
[ 9.53884200e-03, 0.00000000e+00, 0.00000000e+00, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00],
[ 8.22157246e-03, 0.00000000e+00, 0.00000000e+00, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00],
...,
[ 2.30122164e-04, 0.00000000e+00, 0.00000000e+00, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00],
[ 5.40662112e-04, 0.00000000e+00, 0.00000000e+00, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00],
[ 2.90977780e-04, 0.00000000e+00, 0.00000000e+00, ...,
0.00000000e+00, 0.00000000e+00, 0.00000000e+00]],
[[ 2.90977780e-04, 0.00000000e+00, 0.00000000e+00, ...,
-2.01285531e-14, -5.79463896e-11, 2.30793494e-17],
[ 9.53884200e-03, 0.00000000e+00, 0.00000000e+00, ...,
-3.95739321e-13, -1.89960022e-09, 7.56587900e-16],
[ 8.22157246e-03, 0.00000000e+00, 0.00000000e+00, ...,
-1.98813085e-13, -1.63727430e-09, 6.52106644e-16],
...
1.92070767e-07, -5.82489819e-10, 2.18970406e-09],
[ 5.40356997e-04, 1.99644086e-07, 5.15788552e-08, ...,
4.12741922e-07, -1.14739043e-09, 4.69286751e-09],
[ 2.91009975e-04, 2.46249542e-08, 5.26512628e-09, ...,
1.66544953e-07, -3.47304600e-10, 1.97236720e-09]],
[[ 2.91066612e-04, 2.99523228e-08, -2.59348021e-09, ...,
-1.66718407e-07, -3.48121939e-10, 1.77222676e-09],
[ 9.54197122e-03, 1.01752190e-06, -1.14974508e-07, ...,
-9.11224032e-06, -3.17257246e-08, 1.00073073e-07],
[ 8.22501861e-03, 1.05059828e-06, -8.63273167e-08, ...,
-1.34707937e-05, -8.04639253e-08, 1.95221527e-07],
...,
[ 2.29222925e-04, 5.26935391e-07, 5.61885394e-08, ...,
1.92369176e-07, -5.83396090e-10, 2.03285350e-09],
[ 5.40272560e-04, 3.13699724e-07, 2.23127567e-08, ...,
4.12813932e-07, -1.14758955e-09, 4.31984715e-09],
[ 2.91035094e-04, 4.34481998e-08, -1.21067812e-09, ...,
1.66516368e-07, -3.47244310e-10, 1.77041703e-09]]],
shape=(48, 77, 774))
Coordinates:
* Time (Time) float64 384B 0.0 1.999e-14 ... 8.079e-07 1.161e-06
* Atomic velocity (Atomic velocity) float64 616B -2.5 -1.5 ... 1.875 2.5
* Density matrix (Density matrix) <U68 211kB 'ρ<sub>Re, (3S<sub>1/2</sub>...Plot the return flux for each transition:
model.total_flux(sol).visualize()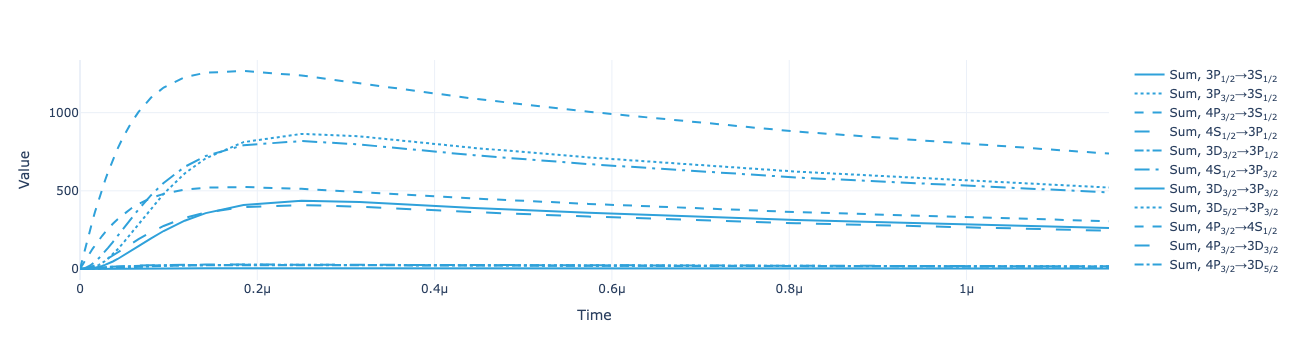
model.total_flux(sol).visualize()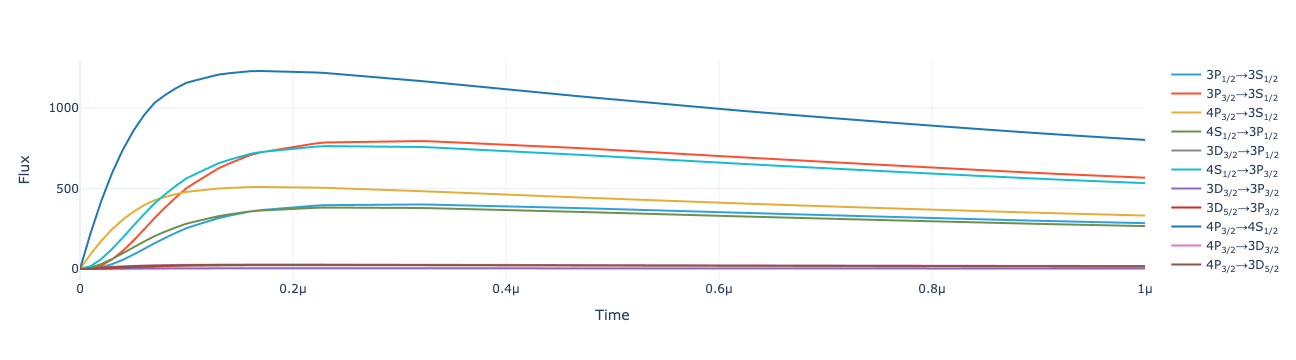
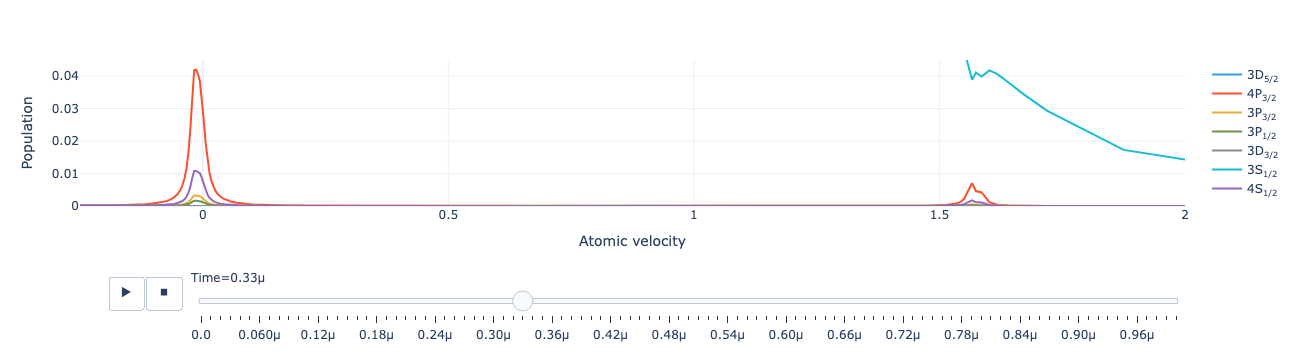
Animate the level population distribution as a function of time:
model.level_population_distribution(sol).visualize(xaxis_range=(-.25, 2), yaxis_range=(0, .045))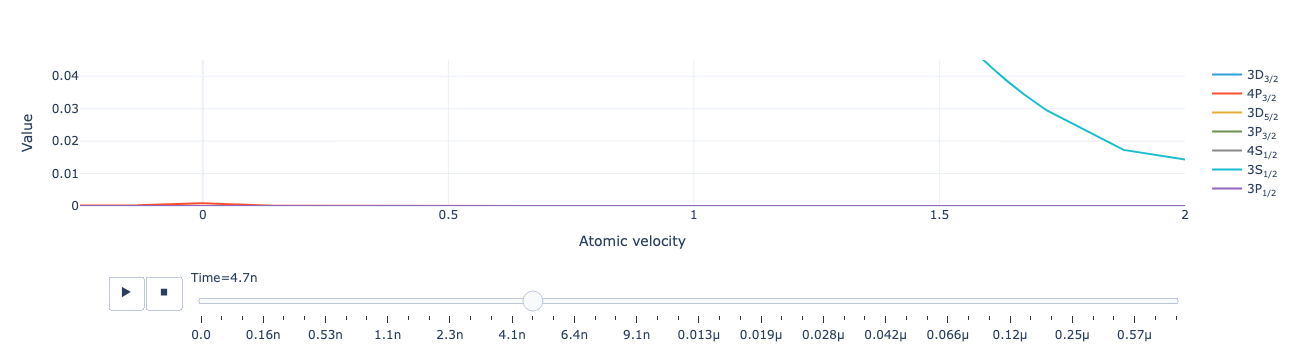
model.level_population_distribution(sol).visualize(xaxis_range=(-.25, 2), yaxis_range=(0, .045))